Lesson 0: Configuring your computer¶
In this lesson, you will set up a Python computing environment for scientific computing. There are two main ways people set up Python for scientific computing on their own machine.
By downloading and installing package by package with tools like apt-get, pip, etc.
By downloading and installing a Python distribution that contains binaries of many of the scientific packages needed. The major distributions of these are Anaconda and Enthought Canopy. Both contain IDEs.
In this class, we will use Anaconda, with its associated package manager, conda. It has become the de facto package manager/distribution for scientific use.
Why install on my own machine?¶
We toyed with using Google Colab for the bootcamp, and we may in fact encourage you to use it when you are working on exercises together remotely in these days of COVID-19. This offers many advantages.
You get a dedicated powerful machine that is part of Google’s cloud resources for your computing.
Most of what you need is pre-installed.
You can easily share the notebook with collaborators in near-real time.
It is free!
There are many more advanced features that are not covered in the bootcamp, such as easy GPU computing.
We chose not to do this for several reasons, the first being the most important.
It is important that you learn how to set up your own machine, including using the command line.
Unless you have a pro account (not free), your Colab instance will shut down after a period of latency.
It is not as easy to integrate into your own software development pipeline.
Along those same lines, it is not as easy to install custom software, or software that you write, using Colab as it is on your own machine.
You may also want to use other cloud computing resources like AWS, Microsoft Azure, or Caltech’s HPC. We are not using those because we do not need more computing power than is available on a standard laptop. Furthermore, setting up your own machine will help prepare you for setting up instances on those services.
Before we get rolling with the Anaconda distribution on your own machine, we have some considerations and installations to get out of the way first.
macOS users: Install XCode¶
If you are using macOS, you should install XCode, if you haven’t already. It’s a large piece of software, taking up about 5GB on your hard drive, so make sure you have enough space. Please install this ahead of the bootcamp, since if students install this during the bootcamp, it will overwhelm the wireless. You can install it through the App Store.
After installing it, you need to open the program. Be sure to do that, for example by clicking on the XCode icon in your Applications folder. Upon opening XCode, it may perform more installations.
Windows users: Install Git and Chrome or Firefox¶
We will be using JupyterLab in the bootcamp. It is browser-based, and Chrome, Firefox, and Safari are supported. Internet Explorer is not. Therefore, if you are a Windows user, you need to be sure you have either Chrome of Firefox installed.
Git is installed on Macs with XCode. For Windows users, you need to install Git. You can do this by following the instructions here.
Uninstalling Anaconda¶
If you have previously installed Anaconda with a version of Python other than 3.7, you need to uninstall it, removing it completely from your computer. You can find instructions on how to do that from the official uninstallation documentation.
Downloading and installing Anaconda¶
Downloading and installing Anaconda is simple.
Go to the Anaconda distribution homepage, click “Download” and download the graphical installer. Be sure to download Anaconda for Python 3.7 for the appropriate operating system.
Follow the on-screen instructions for installation. When prompted, be sure to “Install for me only.”
You may be prompted for optional installations, like PyCharm. You will not need these for the bootcamp.
That’s it! After you do that, you will have a functioning Python distribution.
Launching JupyterLab and a terminal¶
After installing the Anaconda distribution, you should be able to launch the Anaconda Navigator. If you’re using macOS, this is available in your Applications menu. If you are using Windows, you can do this from the Start menu. Launch Anaconda Navigator.
We will be using JupyterLab throughout the bootcamp (more on that in Lesson 1). You should see an option to launch JupyterLab. When you do that, a new browser window or tab will open with JupyterLab running. Within the JupyterLab window, you will have the option to launch a notebook, a console, a terminal, or a text editor. We will use all of these during the bootcamp. For the updating and installation of necessary packages, click on Terminal to launch a terminal. You
will get a terminal window (probably black) with a bash prompt. We refer to this text interface in the terminal as the command line.
The conda package manager¶
conda is a package manager for keeping all of your packages up-to-date. It has plenty of functionality beyond our basic usage in class, which you can learn more about by reading the docs. We will primarily be using conda to install and update packages.
conda works from the command line. Now that you know how to get a command line prompt, you can start using conda. The first thing we’ll do is update the packages that came with the Anaconda distribution. To do this, enter the following on the command line:
conda update --all
If anything is out of date, you will be prompted to perform the updates, and press y to continue. (If everything is up to date, you will just see a list of all the installed packages.) They may even be some downgrades. This happens when there are package conflicts where one package requires an earlier version of another. conda is very smart and figures all of this out for you, so you can almost always say “yes” (or “y”) to conda when it prompts you.
Installations¶
There are several additional installations you need to do for the bootcamp. We will first install some plotting packages we need, which are available as HoloViz.
conda install -c pyviz holoviz
Some packages may again be downgraded with the installation of HoloViz, and that is ok. Next, to configure JupyterLab, we need to install node.js. We’ll also install Black, which will make formatting our code easier, and pySerial, which allows for communication with external devices.
conda install nodejs black pyserial
There are a few other packages from pip we will need for the bootcamp, so we can go ahead and install those now.
pip install watermark blackcellmagic bokeh-catplot bootcamp_utils
Finally, we need to configure JupyterLab to work with the plotting packages we will use.
jupyter labextension install --no-build @pyviz/jupyterlab_pyviz
You may also wish to install a spell-checker (this one isn’t necessary).
jupyter labextension install --no-build @ijmbarr/jupyterlab_spellchecker
After installing all of these extensions, you can rebuild JupyterLab.
jupyter lab build
You should close your JupyterLab session and terminate Anaconda Navigator after you have completed the build. Relaunch Anaconda Navigator and launch a fresh JupyterLab instance. As before, after JupyterLab launches, launch a new terminal window so that you can proceed with setting up Git.
Setting up Git¶
Git features heavily in the bootcamp. We use it for version control and also for sharing files, data, and code we will use in the bootcamp.
Set up a GitHub account¶
Go to http://github.com/ to get an account. You should register with your academic email address so you get free private repositories as academics. You should also think carefully about picking your user name. There is a good chance other people in your professional life will see this.
Forking the bootcamp repository¶
Let’s say you want to do some work on a project with code stored in a repository, but you are not an active collaborator. For example, there could be a useful package a lab at another university put on GitHub for a certain kind of image segmentation that is useful for your research. You want to do something almost exactly like the package does, but need to make some small modifications yourself. You want to clone the repository and add a couple functions and maybe modify one or two they already have, leaving much of the rest of the repository untouched. Of course, you also want to update your local copy of all that untouched (but still used) code when the maintainers update it.
This is kind of exactly what you want to do here in the bootcamp. We have a repository that has data sets and a couple tutorials, but you want to write your Python code right in that repository. If I update the data sets, you want to be able to pull in my changes, but still have your code in place.
There is a nice way to do this called forking. To fork a repository on GitHub, simply navigate got the website of the repository and click the Fork button. Be sure you are logged in as yourself when you do this. Click this link to go to the GitHub page for the bootcamp.
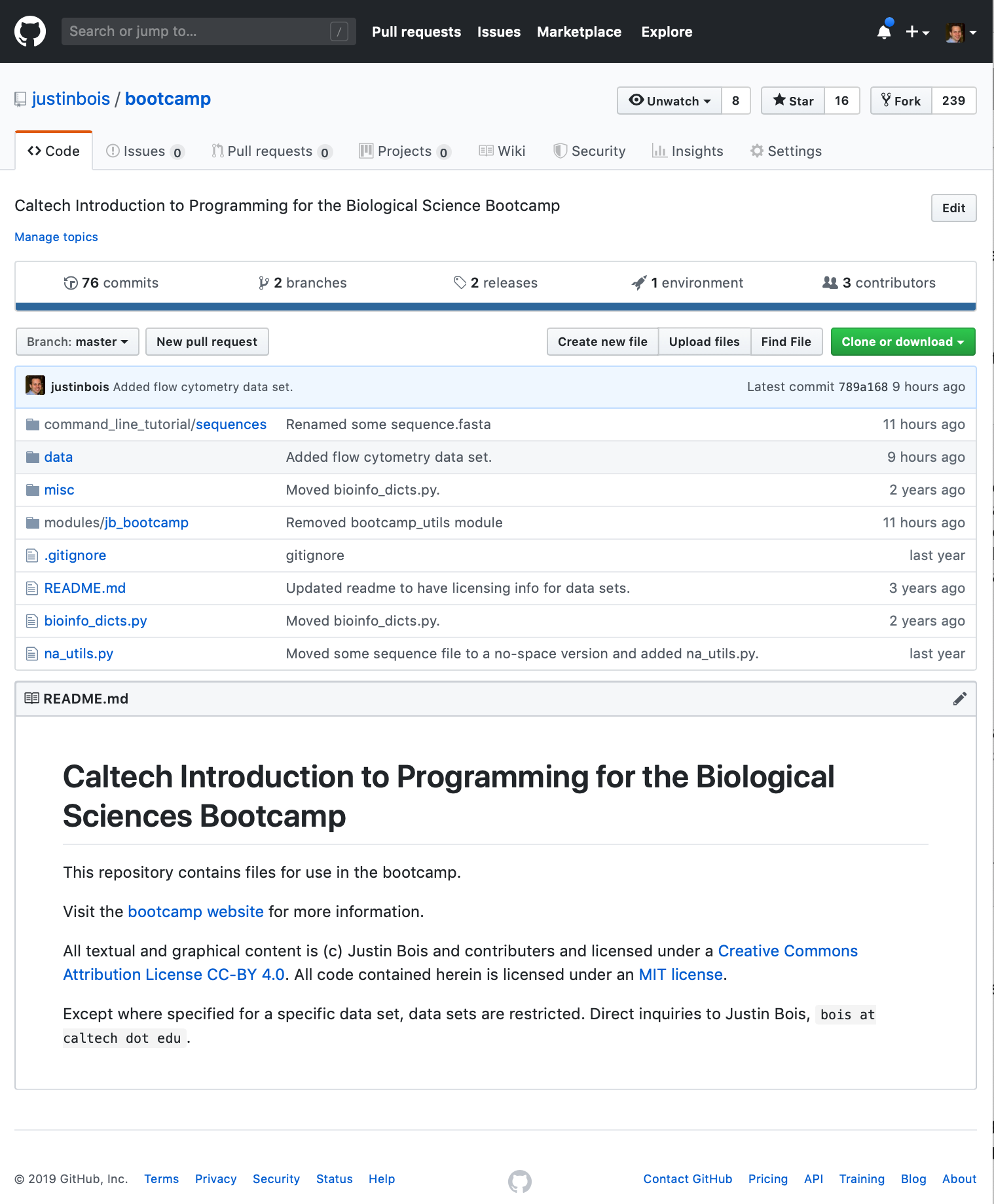
The fork button is in the upper right. Just click the button, and you now have a fork of the bootcamp repository on your GitHub account.
Cloning your fork to your local machine¶
Now you can clone your fork of the repository to your local machine. We will keep all of your material under version control in a directory called git in your home directory. Do the following on the command line.
mkdir -p ~/git
Now you can clone your forked copy of the repository (not the original bootcamp repository). To do this, navigate your browser to the forked copy of the bootcamp repository on your account (this is where clicking the “Fork” button took you in your browser). The browser URL will be: https://github.com/yourusername/bootcamp, and the top left of the website will say “yourusername/bootcamp forked from justinbois/bootcamp, like below for user cgoecknerwald.
![]()
Click the green Clone or download button (shown below) and copy the URL of the forked repository.
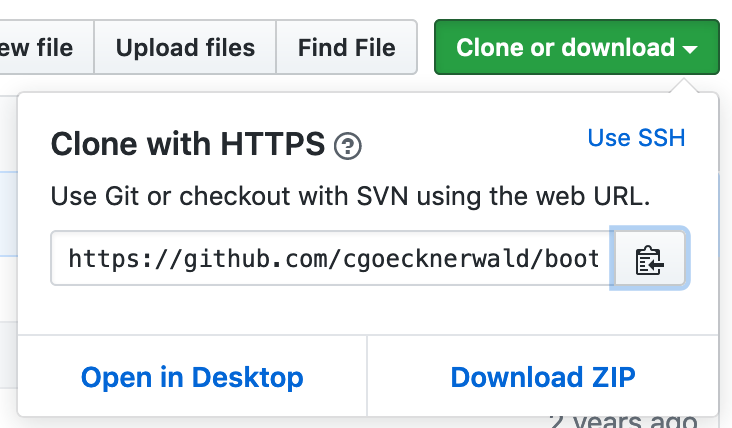
Now, you can clone it by doing the following on the command line, making the obvious substitution for the_url_you_just_copied.
cd ~/git
git clone --depth 5 the_url_you_just_copied
You now have a local copy of your own fork of the bootcamp repository. You can add files and edit it. When you commit and push, it will all be on your account, and the master repository will not see the changes. (Don’t worry if you do not understand all of this now; we will cover it all in subsequent lessons.)
Syncing your forked repository to the upstream repository¶
As I mentioned before, you want to be able to sync your repository with the original bootcamp repository so you can retrieve any updates in it. The original repository is typically called the upstream repository, since presumably you are changing it, so you are downstream. You want the upstream repository to be a remote repository, which is just what we call a repository we track and fetch and merge from. To see which repositories are remote, do the following on the command line.
cd bootcamp
git remote -v
The -v just means “verbose,” so it will also tell you the URLs. Entering that now will show a single repository, origin, which you can fetch from and push to. In your case, origin your fork of the bootcamp repository.
We now want to add the upstream repository. To do this, add the original bootcamp repository as the upstream repository.
git remote add upstream https://github.com/justinbois/bootcamp.git
Now try doing git remote -v, and you will see that you are now also tracking the upstream repository.
Checking your distribution¶
We’ll now run a quick test to make sure things are working properly. We will make a quick plot that requires some of the scientific libraries we will use in the bootcamp.
Use the JupyterLab launcher (you can get a new launcher by clicking on the + icon on the left pane of your JupyterLab window) to launch a notebook. In the first cell (the box next to the [ ]: prompt), paste the code below. To run the code, press Shift+Enter while the cursor is active inside the cell. You should see a plot that looks like the one below. If you do, you have a functioning Python environment for scientific computing!
[1]:
import numpy as np
import bokeh.plotting
import bokeh.io
bokeh.io.output_notebook()
# Generate plotting values
t = np.linspace(0, 2*np.pi, 200)
x = 16 * np.sin(t)**3
y = 13 * np.cos(t) - 5 * np.cos(2*t) - 2 * np.cos(3*t) - np.cos(4*t)
p = bokeh.plotting.figure(height=250, width=275)
p.line(x, y, color='red', line_width=3)
text = bokeh.models.Label(x=0, y=0, text='bootcamp', text_align='center')
p.add_layout(text)
bokeh.io.show(p)
Computing environment¶
[2]:
%load_ext watermark
%watermark -v -p numpy,bokeh,jupyterlab
CPython 3.7.7
IPython 7.13.0
numpy 1.18.1
bokeh 2.0.2
jupyterlab 1.2.6